Data Stewardship
Due to the current circumstances there are now face to face RDM counseling hours. If you have questions please write an email to us and we will arrange a virtual meeting.
UBC Research Data Management Workshop Series 2020
In the last quarter of 2019 we have offered a RDM specific workshop series for the first time, you can find the flyer here: UBC-RDM-WorkshopSeries-2019
We have received great feedback and had good attendance numbers throughout the series. We are currently working on creating the workshop schedule for this year and will inform you in time when to sign up!
This workshop series is open for everyone (from masters students to PI-level, and staff of course) from the members of the UBC and ONCODE associates and X-Omics colleagues.
You need some quick pointers and help how to improve your personal research data management? Then you are on the right page. The following 8 cues will provide you with a solid start into good research data management practises:
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
- Create a coherent folder and file structure
- Establish File Naming Conventions
- Improve the Study Documentation
- Create a regular Back-Up Routine
- Get a free ORCID-ID
- Publish data on a platform that hands out persistent links
- Cite your data-set properly
- Rely on ALTMETRICS and Co.
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
Click on the grey dots to get to the next explanation in the slider below.
1. Create a coherent folder and file system
Step 1: Create a coherent folder and file system
Step 2: Establish version control
Step 3: Write readme files
-
-
-
-
-
-
-
-
-
- Creating a logical folder structure with a few top-level folders to divide between research and administrative content, and establishing a number of sub-folders at the beginning of a new project helps with managing, sharing, and saving files over time.
- Adding versioning numbers on higher level folders, and all files helps keeping the overview and statuses of their contents independent from time-stamps in the file-directories.
- Finally, adding simple text-files with a minimum set of information (aka readme file) about all subordinate folders and files for each top-level folder will increase the comprehensibility for you and your colleagues in the course of the project and beyond.
-
-
-
-
-
-
-
-
Read more about file and folder management: https://libraries.mit.edu/data-management/files/2014/05/FileOrg_20160121.pdf
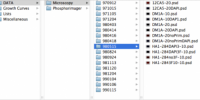
2. Establish File Naming Conventions
Step 1: Start adding the last modification date in a standardised Syntax in every new file and folder title
Step 2: Add Version numbers in in every new file title
Step 3: Create and add abbreviations and identifiers to every new file and folder title
1. From now on, add the last modification date to every new file title you write. The international standard to write a date is YYYY-MM-DD (year, month, date), out of convenience the dash can be left out.
Example: Test_190517
2. Additionally, adding a version number with integers will quickly reveal what the latest version of a file is.
Example: Test_V5_190518
3. Creating abbreviations and identifiers for processes or applied methodologies enhances the quick comprehension of the file contents in the directory without the need to open the actual file.
Example: <analysis_name>.filtered_variants.snpEff_GoNLv5.vcf
Read more about File Naming Conventions: http://www.lse.ac.uk/library/research-support/research-data-management/version-control
3. Improve the Study Documentation
Step 1: Write about the relationship and context between files within the folder and file structure.
Step 2: Add information about vocabulary, terminology and definitions that you have applied.
Step 3: Continue with explaining the chosen methodology.
-
-
-
-
-
-
-
-
-
-
- For starters, by offering some contextual information how files and folders within a project directory relate to each other, users of these files can grasp their relationship to each other more easily.
- By adding explanation to applied vocabulary, terms used, and domain specific definitions, the framework in which your research happened is more transparent and reproducible.
- Lastly, specifying which methodologies were applied for which data-files will complete the picture of your research that will enable authentic reusability.
-
-
-
-
-
-
-
-
-
Read more about documentation writing: https://weblog.wur.eu/openscience/documenting-research-data-along-way-tips-tools/
4. Create a regular Back-Up Routine
Step 1: Decide and adhere to a regular back-up habit.
Step 2: Consider automising this regular back-up routine.
Step 3: Store your back-up data ideally in 3 different locations.
-
-
-
-
-
-
-
-
-
-
- Depending on the uniqueness of your data and the sensitively level, the intervals of transferring your important files to a back-up location might be different. At least a weekly back-up is a good start.
- By applying back-up management software, the effort of taking care of regular back-ups can be reduced.
- Ideally, your regular back-up will be spread across different and independent and secure storage locations. Besides your working computer, institutionally provided storage drives are a good option for another back-up location.
-
-
-
-
-
-
-
-
-
Do you know about SurfDrive? To all members of the UBC SurfDrive is accessible right away; it offers an online interface and a desktop-client with inter-device synchronisation.
Read more about SurfDrive: https://www.surf.nl/en/store-and-share-your-files-securely-in-the-cloud-with-surfdrive
5. Get a free ORCID-ID
Image reference: https://orcid.org/
Sign up for your freely available unique authors ID. In addition to most of the publishers these days requiring this ID-type already,
the ORCID-ID helps you being individually identified as creator and will list all outputs that is related to this ID.
Publishing other scientific material such as poster, presentation slides, graphics, software, and videos will also count as publication
and will accumulate in your publishing record.
Read more about the ORCID-ID and register: https://orcid.org/register
6. Publish data on a platform that hands out persistent links
Image reference: https://commons.wikimedia.org/wiki/File:DOI_logo.svg
Reconsider sharing finished data-sets and other files via email upon request, or placing it as individual file on your project website:
rather self-publish it on a general data repository such as Zenodo, to receive a persistent link and therefore all benefits of citations and discoverability with it.
Another advantage: you can make the publication underlying data available as its own entity with a DOI.
Set-up properly, both the publication and the data-set will have their own persistent links and refer to each other.
That can possibly lead to a scenario, where your data is independently cited and re-used as much and perhaps more than the related publication.
Read more about how Zenodo enables DOI versioning: https://help.zenodo.org/
7. Cite your data-set properly
Image reference: https://www.flaticon.com/free-icon/network_148800
The best way to cite your own data is to refer to it in-text (not in the footnote!) or in the reference section. That way it can be detected and picked up by citation tools used by publishers and library indices that feed into your citation score. Instead of sharing your data files via a website with a URL that might be unstable over time, choose to publish your data via a dedicated repository or archive that will attach a persistent link to that data-set. Any type of publication with a persistent link will accumulate more citations over time due to accessibility and discoverability on the web.
Read more on data citation styles: http://libraryguides.vu.edu.au/c.php?g=386501&p=4347840
8. ALTMETRICS and CO.
Image reference: https://www.altmetric.com/about-us/logos/
The way science is conducted and documented is changing, so is the academic reward system. A future is on the horizon where
data and software publications, as well a plethora of other output forms of scientific work next to the classic and prestigious publications
are recognised as valuable and impactful body of works. One popular employed by many universities across the globe is
called ALMETRICS. Besides the classic citation tracking it enables the attention and impact detection and tracking of all sorts of outputs
ranging from tweets, over opinions in comment sections, to policy papers. The impact of your work on the wider society can now
be traced and real time interaction with peers facilitated. Due to ORCID-ID’s and DOIS this transparency and historical hashing
of interactions is possible.
Read more about ALTMETRICS: https://www.altmetric.com/audience/researchers/
These helpful cues picked your brain and now you want to know more, or have any other matter you would like to discuss?
Please drop me a few lines via email using the contact button below.
Looking forward to your message!
Jasmin K. Böhmer, MSc | Data Steward | Center for Molecular Medicine and Oncode Institute | UBC Utrecht Data Stewardship